ClearLight Technology
CLARITY Tissue Clearing and 3D IHC
ClearLight empowers researchers to see more biology.
Tru3D™* COMPARISON
The Problem
Current gold standard techniques are 100 years old and rely on the analysis of 2D thin section FFPE (~5-10 micron). Spatial and morphological analysis of the tissue microenvironment is highly limited. 2D is destructive, slide-based, non-digital, is variable, and highly manual and does not represent the entire biology.
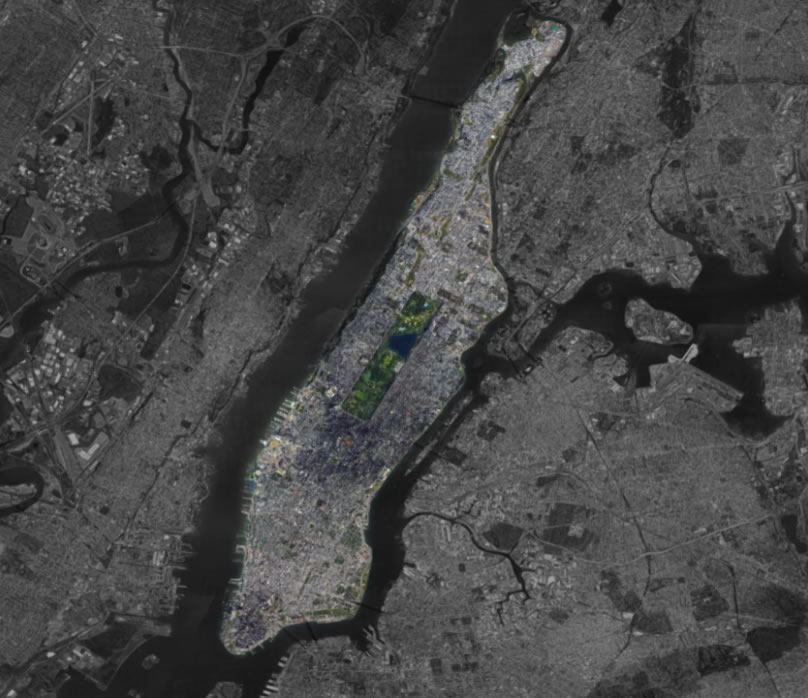
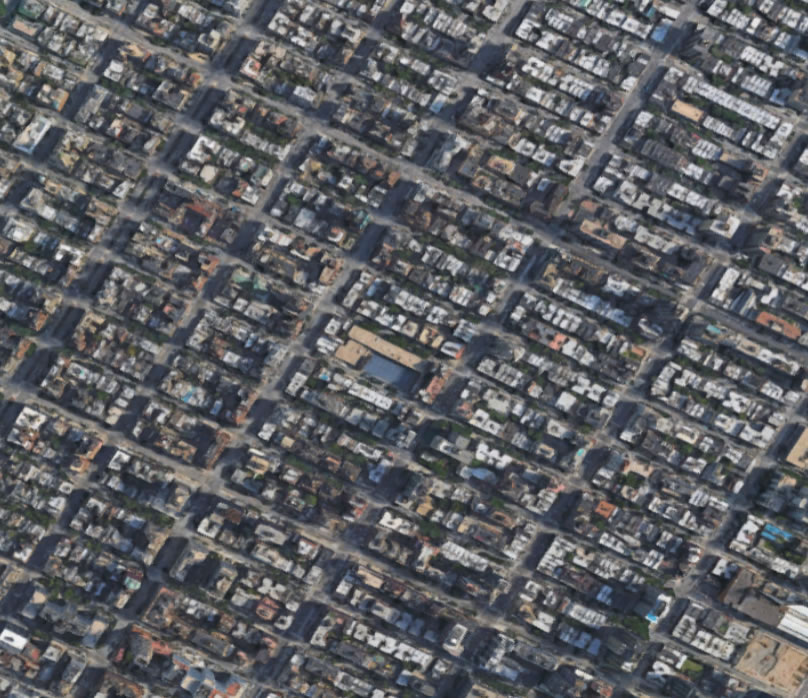
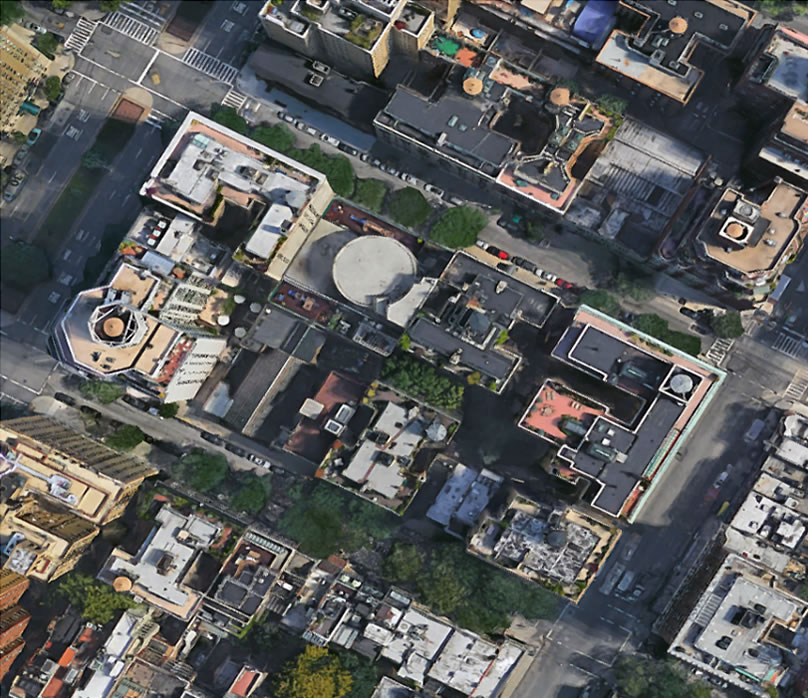
A Single City Block vs. All of Manhattan
-
Mathematically speaking...

-
Mathematically speaking…
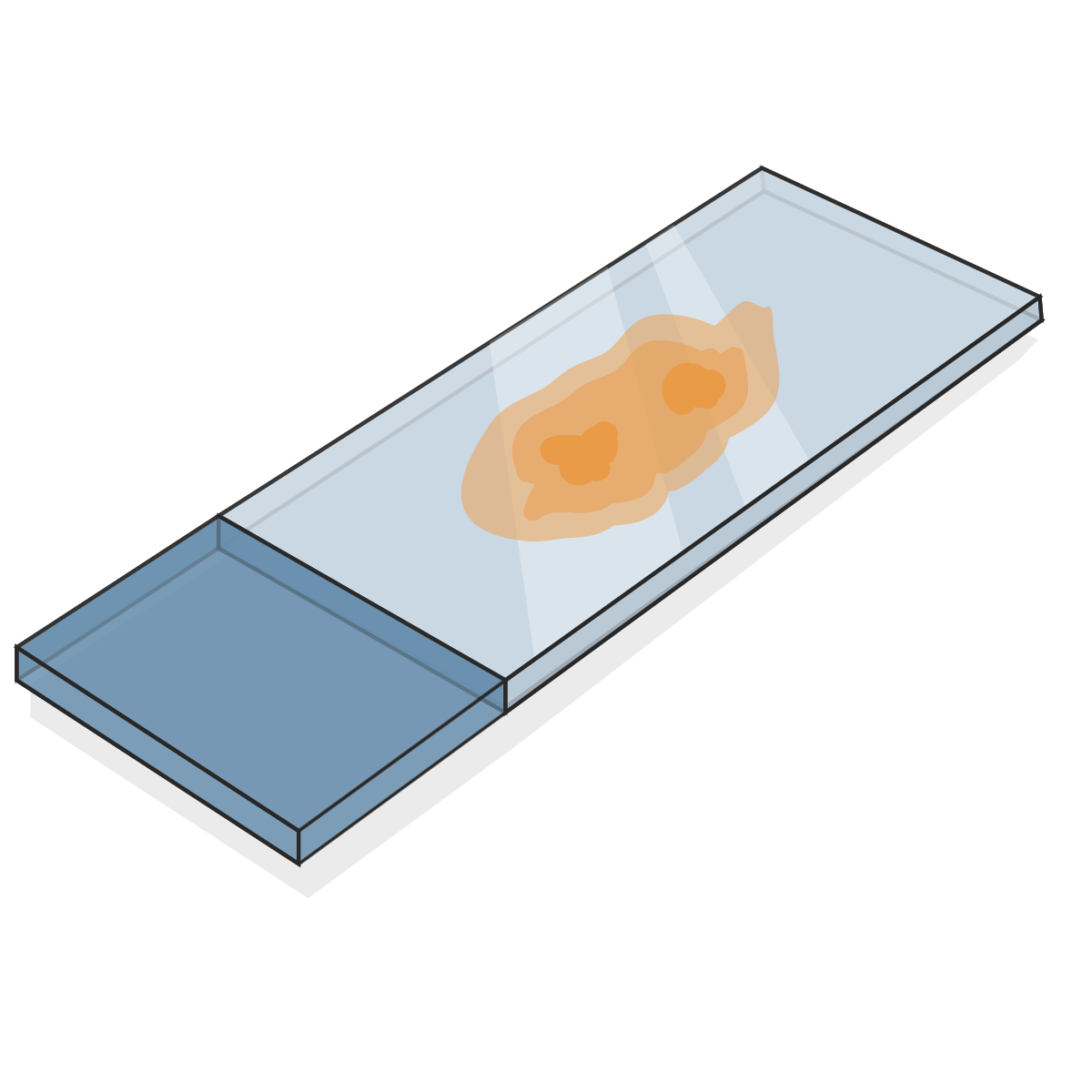
A single, 5µm tissue section from a 2 cm3 breast carcinoma,
-
Mathematically speaking…
A single, 5µm tissue section from a 2 cm3 breast carcinoma,
is exactly the same as the ratio of a single Manhattan block to all of Manhattan

-
Mathematically speaking…
A single, 5µm tissue section from a 2 cm3 breast carcinoma,
is exactly the same as the ratio of a single Manhattan block to all of Manhattan
Adapted with permission from Geoffrey Baird MD, PhD
Not representative
of the whole!

The Solution
See more biology in Tru3D. Bring CLARITY technology to your tissue processing workflow. Use 3D histopathology in your research.
CLARITY Overview
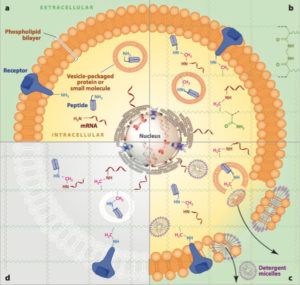
CLARITY is an acronym for Clear Lipid-exchanged Acrylamide-hybridized Rigid Imaging / Immunostaining / in situ-hybridization-compatible Tissue hYdrogel)
Tissue Processing Workflow using CLARITY technology
CLARITY tissue clearing embeds tissue in a hydrogel matrix. Lipids are cleared to create an optically transparent hybrid that is transparent to light and permeable to molecules. Scientists can then add molecular markers to highlight specific features for further imaging and analysis. The tissue processing workflow can also include washing out staining, allowing for multiple rounds of molecular labeling and imaging.
Benefits of using CLARITY in Tissue Processing Workflow
- Compatible with previously frozen, fresh, formalin-fixed, and FFPE embedded tissue
- Compatible with traditional pathology workflow methods
- Non-destructive, fewer samples, allowing recapitulation of spatial heterogeneity
- No dehydration/rehydration steps involved as with FFPE samples, or other tissue clearing technologies
- Compatible with standard nucleic acid and antibody interrogation techniques
- Samples can be stored for future biomarker analysis
- Imaging compatibilities include: Confocal, Light Sheet, SPIM
CLARITY Tissue Processing Workflow
Mouse, Rat, Human
Fixed • Fresh • Frozen • FFPE
Hydrogel Matrix
The first step in the tissue processing workflow is to place the mouse, rat, or human tissue sample in a solution of hydrogel monomers and cross-linkers. For detailed explanations refer to Hydrogel-Tissue Chemistry.
Lipid Clearing
A detergent is used to wash lipids and other unbound molecules from the tissue. The proteins, nucleic acids and other bound biomolecules remain embedded with the hydrogel mesh.
Immunostaining
If desired, antibody-based immunostaining or labeling for many nucleic acids (RNA/DNA) at once can be used to highlight specific structures in the clarified sample.
Image Analysis
The tissue processing workflow moves from molecular labeling to imaging. Learn more about about immunohistochemistry (3D IHC). The stained tissue is placed in a mounting solution for imaging with a confocal or light sheet microscope or another 3D technique.
Tumor Microenvironment
A frozen human stage IB squamous cell carcinoma lung cancer tissue was cleared using CLARITY and imaged. Tissue autofluorescence demonstrates intact tissue morphology of this lung cancer specimen.
Imaging Comparison
Tru3D overcomes the limitations of 2D imaging, which is not representative of the heterogeneous biopsy or tissue microenvironment. See a 2D Versus Tru3D comparison of tissue processing and image analysis.
Machine Learning
Our artificial intelligence (AI) algorithms are still learning. Your research team can benefit from working with our tissue processing and imaging workflow. Together we can explore further and see more biology. Collaborate With Us.

Publications
- Chen Y, Shen Q, White SL, Gokmen-Polar Y, Badve S, & Goodman LJ. Three-dimensional imaging and
quantitative analysis in CLARITY processed breast cancer tissues. Nature: Scientific Reports, (2019)
9:5624. - Wang X, Allen WE, Wright MA, Sylwestrak EL, Samusik N, Vesuna S, Evans K, Liu C, Ramakrishnan C, Liu J, Nolan GP, Bava FA, Deisseroth K. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science. 2018 Jul 27;361(6400)
- Gradinaru V, Treweek J, Overton K, Deisseroth K. Hydrogel-tissue chemistry: principles and applications. Annual Review of Biophysics, 2018 May 20;47:355-376.
- Hsueh B, Burns VM, Pauerstein P, Holzem K, Ye L, Engberg K, Wang A-C, Gu X, Chakravarthy H, Arda HE, Charville G, Vogel H, E mov IR, Kim S, Deisseroth K. Pathways to clinical CLARITY: volumetric analysis of irregular, soft, and heterogeneous tissues in development and disease. Sci Rep. 2017 Jul 19;7(1):5899.
- Deisseroth K. A look inside the brain. Scientific American, 2016 Sep 20;315(4):30-37
- Sylwestrak EL, Rajasethupathy P, Wright MA, Jaffe A, Deisseroth K. Multiplexed intact-tissue transcriptional analysis at cellular resolution. Cell. 2016 Feb 11;164(4):792-804
- Chung K, Deisseroth K. CLARITY for mapping the nervous system. Nature Methods. June 2013. 10(6):508-13.
- Structural and molecular interrogation of intact biological systems. Chung K, Wallace J, Kim SY, Kalyanasundaram S, Andalman AS, Davidson TJ, Mirzabekov JJ, Zalocusky KA, Mattis J, Denisin AJ, Pak S, Bernstein H, Ramakrishnan C, Grosenick L, Gradinaru V, and K Deisseroth. Nature 497, 332–337 (16 May 2013).
More Technology
Posters
Abstract #4690: Sharla L. White, Yi Chen, Qi Shen, Laurie J. Goodman. Multi-sample automation of the
CLARITY technology for the processing of 3D volumes of tissue. AACR Annual Meeting, March 29-April 3,
2019, Atlanta, GA. [Download] [PDF]
Abstract #351: Three-dimensional (3D) Imaging of Biomarkers in Human Core Needle Biopsies of Normal and Cancerous Breast Tissue. Yi Chen, Qi Shen, Laurie J. Goodman, Yesim Gokmen-Polar, Sunil Badve. SABCS Annual Meeting, Dec 4-8, 2018, San Antonio, TX. [Download] [PDF]
Abstract 5915: Three-dimensional, 3-D, multiplex imaging of biomarkers in tumor tissue. Sharla L. White, Sarah McCurdy and Laurie J. Goodman. AACR Annual Meeting 2017; April 1-5, 2017; Washington, DC [Download] [PDF]